You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004461_03370
You are here: Home > Sequence: MGYG000004461_03370
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Verrucomicrobiota; Lentisphaeria; Victivallales; UBA1829; ; | |||||||||||
| CAZyme ID | MGYG000004461_03370 | |||||||||||
| CAZy Family | GH36 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 24464; End: 26605 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH36 | 93 | 648 | 7.3e-88 | 0.7805232558139535 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| cd14791 | GH36 | 1.62e-75 | 324 | 616 | 1 | 299 | glycosyl hydrolase family 36 (GH36). GH36 enzymes occur in prokaryotes, eukaryotes, and archaea with a wide range of hydrolytic activities, including alpha-galactosidase, alpha-N-acetylgalactosaminidase, stachyose synthase, and raffinose synthase. All GH36 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH36 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
| pfam02065 | Melibiase | 1.05e-34 | 318 | 624 | 34 | 347 | Melibiase. Glycoside hydrolase families GH27, GH31 and GH36 form the glycoside hydrolase clan GH-D. Glycoside hydrolase family 36 can be split into 11 families, GH36A to GH36K. This family includes enzymes from GH36A-B and GH36D-K and from GH27. |
| COG3345 | GalA | 1.77e-25 | 99 | 531 | 83 | 496 | Alpha-galactosidase [Carbohydrate transport and metabolism]. |
| cd14792 | GH27 | 9.93e-06 | 329 | 535 | 5 | 174 | glycosyl hydrolase family 27 (GH27). GH27 enzymes occur in eukaryotes, prokaryotes, and archaea with a wide range of hydrolytic activities, including alpha-glucosidase (glucoamylase and sucrase-isomaltase), alpha-N-acetylgalactosaminidase, and 3-alpha-isomalto-dextranase. All GH27 enzymes cleave a terminal carbohydrate moiety from a substrate that varies considerably in size, depending on the enzyme, and may be either a starch or a glycoprotein. GH27 members are retaining enzymes that cleave their substrates via an acid/base-catalyzed, double-displacement mechanism involving a covalent glycosyl-enzyme intermediate. Two aspartic acid residues have been identified as the catalytic nucleophile and the acid/base, respectively. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ANS77143.1 | 5.89e-168 | 68 | 711 | 34 | 686 |
| BBF45383.1 | 5.57e-166 | 5 | 713 | 3 | 700 |
| BBI32765.1 | 5.71e-166 | 8 | 711 | 18 | 710 |
| QYY36366.1 | 1.24e-164 | 69 | 713 | 57 | 711 |
| QYM79268.1 | 4.59e-162 | 13 | 713 | 16 | 725 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4FNQ_A | 9.83e-33 | 276 | 702 | 282 | 709 | Crystalstructure of GH36 alpha-galactosidase AgaB from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 3MI6_A | 5.77e-32 | 236 | 651 | 243 | 662 | ChainA, Alpha-galactosidase [Levilactobacillus brevis ATCC 367],3MI6_B Chain B, Alpha-galactosidase [Levilactobacillus brevis ATCC 367],3MI6_C Chain C, Alpha-galactosidase [Levilactobacillus brevis ATCC 367],3MI6_D Chain D, Alpha-galactosidase [Levilactobacillus brevis ATCC 367] |
| 4FNP_A | 9.66e-32 | 276 | 702 | 282 | 709 | Crystalstructure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_B Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_C Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNP_D Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNS_A Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_B Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_C Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus],4FNS_D Crystal structure of GH36 alpha-galactosidase AgaA A355E from Geobacillus stearothermophilus in complex with 1-deoxygalactonojirimycin [Geobacillus stearothermophilus] |
| 4FNR_A | 1.29e-31 | 276 | 702 | 282 | 709 | Crystalstructure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_B Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_C Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus],4FNR_D Crystal structure of GH36 alpha-galactosidase AgaA from Geobacillus stearothermophilus [Geobacillus stearothermophilus] |
| 2XN0_A | 5.40e-31 | 218 | 703 | 227 | 714 | Structureof alpha-galactosidase from Lactobacillus acidophilus NCFM, PtCl4 derivative [Lactobacillus acidophilus NCFM],2XN0_B Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM, PtCl4 derivative [Lactobacillus acidophilus NCFM],2XN1_A Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with TRIS [Lactobacillus acidophilus NCFM],2XN1_B Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with TRIS [Lactobacillus acidophilus NCFM],2XN1_C Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with TRIS [Lactobacillus acidophilus NCFM],2XN1_D Structure of alpha-galactosidase from Lactobacillus acidophilus NCFM with TRIS [Lactobacillus acidophilus NCFM] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| Q9ALJ4 | 7.04e-31 | 276 | 702 | 282 | 709 | Alpha-galactosidase AgaA OS=Geobacillus stearothermophilus OX=1422 GN=agaA PE=1 SV=1 |
| G1UB44 | 2.95e-30 | 218 | 703 | 227 | 714 | Alpha-galactosidase Mel36A OS=Lactobacillus acidophilus (strain ATCC 700396 / NCK56 / N2 / NCFM) OX=272621 GN=melA PE=1 SV=1 |
| P43467 | 1.19e-28 | 208 | 703 | 214 | 713 | Alpha-galactosidase 1 OS=Pediococcus pentosaceus OX=1255 GN=agaR PE=3 SV=1 |
| P27756 | 1.00e-25 | 280 | 524 | 278 | 525 | Alpha-galactosidase OS=Streptococcus mutans serotype c (strain ATCC 700610 / UA159) OX=210007 GN=aga PE=3 SV=3 |
| B8NWY6 | 4.41e-25 | 143 | 527 | 187 | 559 | Probable alpha-galactosidase C OS=Aspergillus flavus (strain ATCC 200026 / FGSC A1120 / IAM 13836 / NRRL 3357 / JCM 12722 / SRRC 167) OX=332952 GN=aglC PE=3 SV=2 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
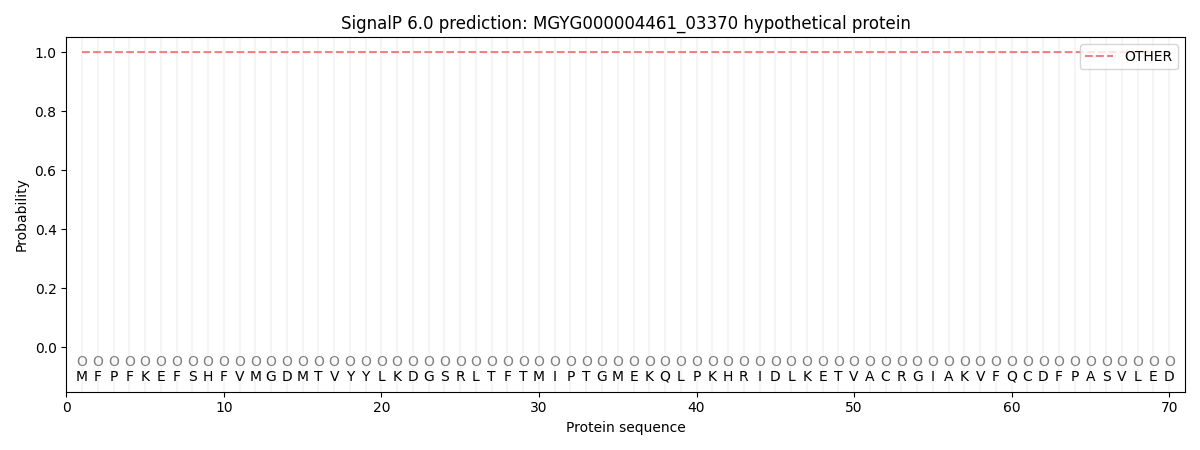
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000069 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
