You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004478_00260
You are here: Home > Sequence: MGYG000004478_00260
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
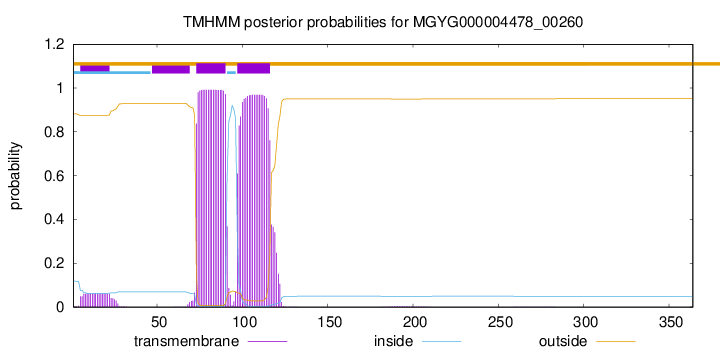
TMHMM annotations
Basic Information help
| Species | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Proteobacteria; Alphaproteobacteria; RF32; CAG-977; UBA11549; | |||||||||||
| CAZyme ID | MGYG000004478_00260 | |||||||||||
| CAZy Family | GT28 | |||||||||||
| CAZyme Description | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 47211; End: 48305 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GT28 | 187 | 349 | 3.7e-41 | 0.9872611464968153 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| PRK00726 | murG | 9.93e-131 | 8 | 363 | 4 | 357 | undecaprenyldiphospho-muramoylpentapeptide beta-N- acetylglucosaminyltransferase; Provisional |
| cd03785 | GT28_MurG | 2.18e-112 | 8 | 355 | 2 | 349 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. MurG (EC 2.4.1.227) is an N-acetylglucosaminyltransferase, the last enzyme involved in the intracellular phase of peptidoglycan biosynthesis. It transfers N-acetyl-D-glucosamine (GlcNAc) from UDP-GlcNAc to the C4 hydroxyl of a lipid-linked N-acetylmuramoyl pentapeptide (NAM). The resulting disaccharide is then transported across the cell membrane, where it is polymerized into NAG-NAM cell-wall repeat structure. MurG belongs to the GT-B structural superfamily of glycoslytransferases, which have characteristic N- and C-terminal domains, each containing a typical Rossmann fold. The two domains have high structural homology despite minimal sequence homology. The large cleft that separates the two domains includes the catalytic center and permits a high degree of flexibility. |
| COG0707 | MurG | 3.70e-101 | 8 | 363 | 3 | 357 | UDP-N-acetylglucosamine:LPS N-acetylglucosamine transferase [Cell wall/membrane/envelope biogenesis]. |
| TIGR01133 | murG | 1.70e-86 | 8 | 357 | 3 | 348 | undecaprenyldiphospho-muramoylpentapeptide beta-N-acetylglucosaminyltransferase. RM 8449890 RT The final step of peptidoglycan subunit assembly in Escherichia coli occurs in the cytoplasm. RA Bupp K, van Heijenoort J. RL J Bacteriol 1993 Mar;175(6):1841-3 [Cell envelope, Biosynthesis and degradation of murein sacculus and peptidoglycan] |
| pfam04101 | Glyco_tran_28_C | 2.55e-37 | 186 | 353 | 1 | 166 | Glycosyltransferase family 28 C-terminal domain. The glycosyltransferase family 28 includes monogalactosyldiacylglycerol synthase (EC 2.4.1.46) and UDP-N-acetylglucosamine transferase (EC 2.4.1.-). Structural analysis suggests the C-terminal domain contains the UDP-GlcNAc binding site. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| ARJ66991.1 | 1.10e-101 | 1 | 354 | 1 | 358 |
| CCQ72804.1 | 1.96e-99 | 8 | 354 | 8 | 358 |
| CUW38580.1 | 2.87e-99 | 8 | 354 | 8 | 358 |
| QQP90937.1 | 5.90e-99 | 3 | 364 | 5 | 369 |
| ACI98048.1 | 1.48e-98 | 4 | 361 | 2 | 362 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 3S2U_A | 1.43e-59 | 8 | 363 | 5 | 357 | Crystalstructure of the Pseudomonas aeruginosa MurG:UDP-GlcNAc substrate complex [Pseudomonas aeruginosa PAO1] |
| 7D1I_A | 4.72e-57 | 8 | 351 | 12 | 356 | ChainA, UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase [Acinetobacter baumannii],7D1I_B Chain B, UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase [Acinetobacter baumannii],7D1I_C Chain C, UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase [Acinetobacter baumannii] |
| 1F0K_A | 1.70e-40 | 1 | 362 | 2 | 355 | The1.9 Angstrom Crystal Structure Of E. Coli Murg [Escherichia coli],1F0K_B The 1.9 Angstrom Crystal Structure Of E. Coli Murg [Escherichia coli],1NLM_A Crystal Structure Of Murg:glcnac Complex [Escherichia coli],1NLM_B Crystal Structure Of Murg:glcnac Complex [Escherichia coli] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| B6IRG2 | 2.96e-99 | 4 | 361 | 2 | 362 | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase OS=Rhodospirillum centenum (strain ATCC 51521 / SW) OX=414684 GN=murG PE=3 SV=1 |
| Q2W0H3 | 1.05e-97 | 8 | 354 | 8 | 358 | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase OS=Magnetospirillum magneticum (strain AMB-1 / ATCC 700264) OX=342108 GN=murG PE=3 SV=1 |
| Q2RVU4 | 2.52e-94 | 6 | 361 | 17 | 373 | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase OS=Rhodospirillum rubrum (strain ATCC 11170 / ATH 1.1.1 / DSM 467 / LMG 4362 / NCIMB 8255 / S1) OX=269796 GN=murG PE=3 SV=1 |
| Q8CY39 | 1.43e-91 | 8 | 354 | 10 | 358 | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase OS=Brucella suis biovar 1 (strain 1330) OX=204722 GN=murG PE=3 SV=1 |
| A5VRH7 | 1.43e-91 | 8 | 354 | 10 | 358 | UDP-N-acetylglucosamine--N-acetylmuramyl-(pentapeptide) pyrophosphoryl-undecaprenol N-acetylglucosamine transferase OS=Brucella ovis (strain ATCC 25840 / 63/290 / NCTC 10512) OX=444178 GN=murG PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
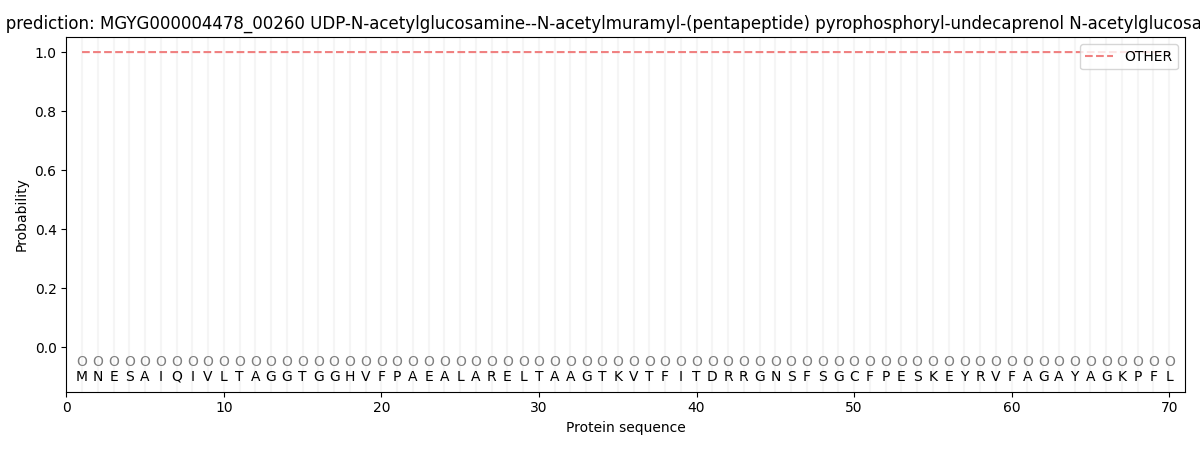
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000074 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |