You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004600_01634
You are here: Home > Sequence: MGYG000004600_01634
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | CAG-307 sp001916215 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Firmicutes; Bacilli; Acholeplasmatales; Anaeroplasmataceae; CAG-307; CAG-307 sp001916215 | |||||||||||
| CAZyme ID | MGYG000004600_01634 | |||||||||||
| CAZy Family | GH105 | |||||||||||
| CAZyme Description | Unsaturated rhamnogalacturonyl hydrolase YteR | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 225; End: 1364 Strand: - | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH105 | 29 | 374 | 7.4e-110 | 0.9789156626506024 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| COG4225 | YesR | 1.71e-87 | 33 | 375 | 31 | 355 | Rhamnogalacturonyl hydrolase YesR [Carbohydrate transport and metabolism]. |
| pfam07470 | Glyco_hydro_88 | 1.89e-73 | 36 | 377 | 27 | 343 | Glycosyl Hydrolase Family 88. Unsaturated glucuronyl hydrolase catalyzes the hydrolytic release of unsaturated glucuronic acids from oligosaccharides (EC:3.2.1.-) produced by the reactions of polysaccharide lyases. |
| COG1331 | YyaL | 0.002 | 29 | 121 | 457 | 569 | Uncharacterized conserved protein YyaL, SSP411 family, contains thoiredoxin and six-hairpin glycosidase-like domains [General function prediction only]. |
| cd04792 | LanM-like | 0.005 | 47 | 129 | 601 | 693 | Cyclases involved in the biosynthesis of class II lantibiotics, and similar proteins. LanM-like proteins. LanM is a bifunctional enzyme, involved in the synthesis of class II lantibiotics. It is responsible for both the dehydration and the cyclization of the precursor-peptide during lantibiotic synthesis. The C-terminal domain shows similarity to LanC, the cyclase component of the lan operon, but the N terminus seems to be unrelated to the dehydratase, LanB. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| CCV66002.1 | 1.16e-180 | 3 | 377 | 2 | 376 |
| CCV64366.1 | 1.54e-166 | 5 | 376 | 6 | 376 |
| CCV64024.1 | 5.46e-165 | 5 | 375 | 7 | 375 |
| ADZ83101.1 | 4.68e-155 | 3 | 373 | 2 | 369 |
| QEH68585.1 | 6.64e-155 | 3 | 373 | 2 | 369 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 4WU0_A | 1.65e-98 | 35 | 372 | 22 | 357 | StructuralAnalysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824],4WU0_B Structural Analysis of C. acetobutylicum ATCC 824 Glycoside Hydrolase From Family 105 [Clostridium acetobutylicum ATCC 824] |
| 1NC5_A | 6.95e-60 | 34 | 374 | 37 | 365 | Structureof Protein of Unknown Function of YteR from Bacillus Subtilis [Bacillus subtilis],2D8L_A Crystal Structure of Unsaturated Rhamnogalacturonyl Hydrolase in complex with dGlcA-GalNAc [Bacillus subtilis] |
| 2GH4_A | 2.94e-59 | 34 | 374 | 27 | 355 | ChainA, Putative glycosyl hydrolase yteR [Bacillus subtilis] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O34559 | 3.80e-59 | 34 | 374 | 37 | 365 | Unsaturated rhamnogalacturonyl hydrolase YteR OS=Bacillus subtilis (strain 168) OX=224308 GN=yteR PE=1 SV=1 |
| P0A3U6 | 1.65e-29 | 186 | 372 | 39 | 227 | Protein Atu3128 OS=Agrobacterium fabrum (strain C58 / ATCC 33970) OX=176299 GN=Atu3128 PE=3 SV=1 |
| P0A3U7 | 1.65e-29 | 186 | 372 | 39 | 227 | 24.9 kDa protein in picA locus OS=Rhizobium radiobacter OX=358 PE=2 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as OTHER
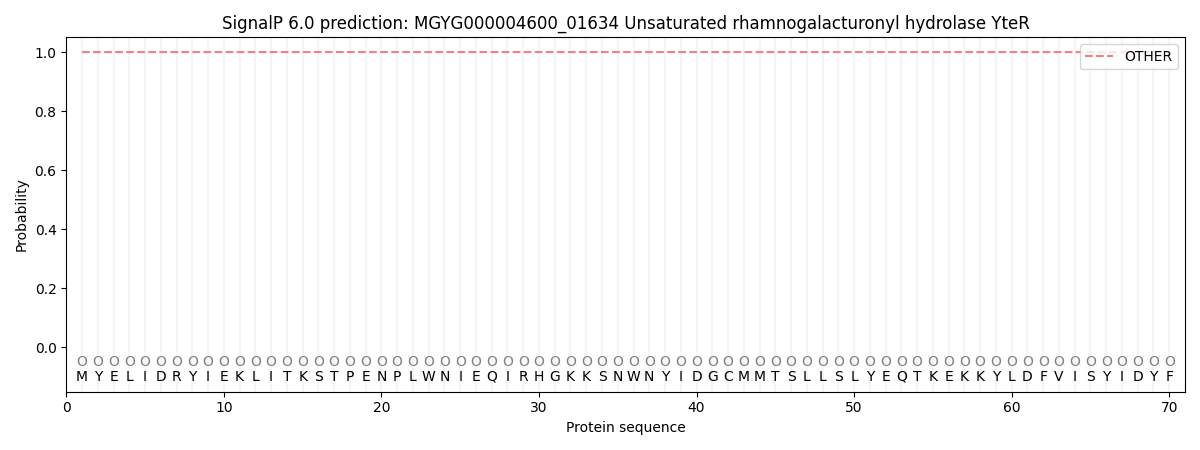
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 1.000039 | 0.000000 | 0.000000 | 0.000000 | 0.000000 | 0.000000 |
