You are browsing environment: HUMAN GUT
MGYG000004872_02479
Basic Information
help
Species
Stoquefichus sp001244545
Lineage
Bacteria; Firmicutes; Bacilli; Erysipelotrichales; Erysipelatoclostridiaceae; Stoquefichus; Stoquefichus sp001244545
CAZyme ID
MGYG000004872_02479
CAZy Family
GH170
CAZyme Description
hypothetical protein
CAZyme Property
Genome Property
Genome Assembly ID
Genome Size
Genome Type
Country
Continent
MGYG000004872
3555940
MAG
Sweden
Europe
Gene Location
Start: 43062;
End: 44213
Strand: -
No EC number prediction in MGYG000004872_02479.
Family
Start
End
Evalue
family coverage
GH170
3
381
3.6e-113
0.9942857142857143
Cdd ID
Domain
E-Value
qStart
qEnd
sStart
sEnd
Domain Description
pfam19200
DUF871_N
2.23e-109
4
240
1
235
DUF871 N-terminal domain. This family consists of several conserved hypothetical proteins from bacteria and archaea. The function of this family is unknown.
more
COG3589
COG3589
1.47e-94
1
383
1
358
Uncharacterized protein [Function unknown].
more
pfam05913
DUF871
5.35e-39
246
381
1
115
Bacterial protein of unknown function (DUF871). This family consists of several conserved hypothetical proteins from bacteria and archaea. The function of this family is unknown.
more
Hit ID
E-Value
Query Start
Query End
Hit Start
Hit End
Description
1X7F_A
8.53e-83
3
373
28
375
Crystalstructure of an uncharacterized B. cereus protein [Bacillus cereus ATCC 14579]
more
Hit ID
E-Value
Query Start
Query End
Hit Start
Hit End
Description
A0A0H2XHV5
1.20e-10
5
383
3
345
6-phospho-N-acetylmuramidase OS=Staphylococcus aureus (strain USA300) OX=367830 GN=mupG PE=1 SV=1
more
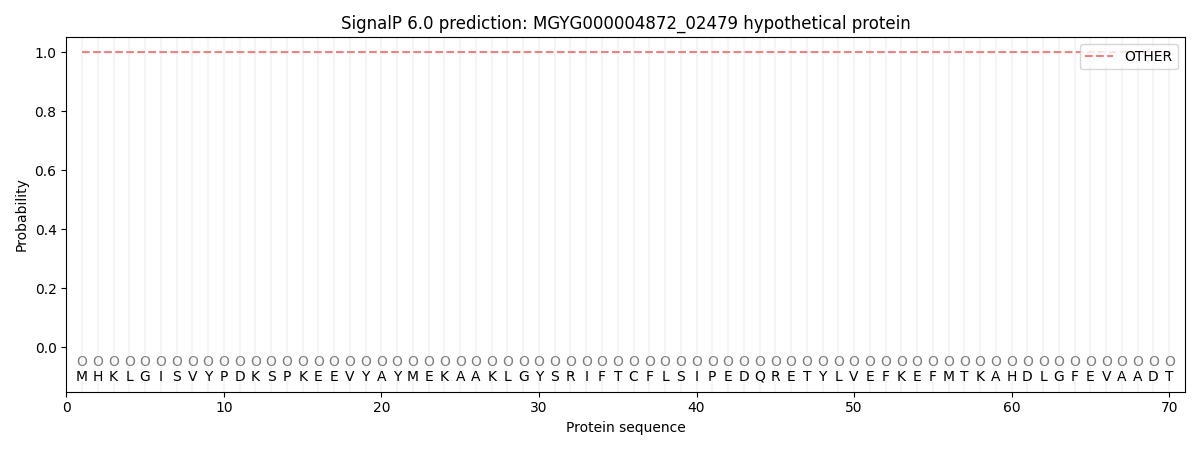
This protein is predicted as OTHER
Other
SP_Sec_SPI
LIPO_Sec_SPII
TAT_Tat_SPI
TATLIP_Sec_SPII
PILIN_Sec_SPIII
1.000064
0.000000
0.000000
0.000000
0.000000
0.000000
There is no transmembrane helices in MGYG000004872_02479.