You are browsing environment: HUMAN GUT
CAZyme Information: MGYG000004899_00736
You are here: Home > Sequence: MGYG000004899_00736
Basic Information |
Genomic context |
Full Sequence |
Enzyme annotations |
CAZy signature domains |
CDD domains |
CAZyme hits |
PDB hits |
Swiss-Prot hits |
SignalP and Lipop annotations |
TMHMM annotations
Basic Information help
| Species | Bacteroides sp900555635 | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Lineage | Bacteria; Bacteroidota; Bacteroidia; Bacteroidales; Bacteroidaceae; Bacteroides; Bacteroides sp900555635 | |||||||||||
| CAZyme ID | MGYG000004899_00736 | |||||||||||
| CAZy Family | GH30 | |||||||||||
| CAZyme Description | hypothetical protein | |||||||||||
| CAZyme Property |
|
|||||||||||
| Genome Property |
|
|||||||||||
| Gene Location | Start: 96041; End: 98200 Strand: + | |||||||||||
CAZyme Signature Domains help
| Family | Start | End | Evalue | family coverage |
|---|---|---|---|---|
| GH30 | 48 | 466 | 3.6e-178 | 0.9928057553956835 |
CDD Domains download full data without filtering help
| Cdd ID | Domain | E-Value | qStart | qEnd | sStart | sEnd | Domain Description |
|---|---|---|---|---|---|---|---|
| pfam02055 | Glyco_hydro_30 | 4.18e-42 | 59 | 406 | 1 | 348 | Glycosyl hydrolase family 30 TIM-barrel domain. |
| COG5520 | XynC | 3.67e-40 | 43 | 470 | 31 | 430 | O-Glycosyl hydrolase [Cell wall/membrane/envelope biogenesis]. |
| pfam17189 | Glyco_hydro_30C | 5.62e-10 | 423 | 466 | 19 | 63 | Glycosyl hydrolase family 30 beta sandwich domain. |
| COG0596 | MhpC | 2.57e-04 | 495 | 718 | 13 | 277 | Pimeloyl-ACP methyl ester carboxylesterase [Coenzyme transport and metabolism, General function prediction only]. |
CAZyme Hits help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End |
|---|---|---|---|---|---|
| QDO67851.1 | 6.35e-316 | 1 | 470 | 1 | 470 |
| ALJ59398.1 | 8.50e-314 | 1 | 469 | 1 | 469 |
| QUT89561.1 | 1.21e-313 | 1 | 469 | 1 | 469 |
| QDO94290.1 | 4.80e-156 | 11 | 466 | 21 | 479 |
| CDF78754.1 | 5.50e-152 | 11 | 469 | 21 | 482 |
PDB Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| 2WNW_A | 1.04e-40 | 56 | 470 | 42 | 444 | Thecrystal structure of SrfJ from salmonella typhimurium [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2],2WNW_B The crystal structure of SrfJ from salmonella typhimurium [Salmonella enterica subsp. enterica serovar Typhimurium str. LT2] |
| 5NGK_A | 5.99e-35 | 42 | 468 | 62 | 490 | Theendo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron],5NGK_B The endo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron],5NGK_C The endo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron],5NGL_A The endo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron],5NGL_B The endo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron],5NGL_C The endo-beta1,6-glucanase BT3312 [Bacteroides thetaiotaomicron] |
| 3KE0_A | 2.79e-34 | 48 | 470 | 67 | 496 | ChainA, Glucosylceramidase [Homo sapiens],3KE0_B Chain B, Glucosylceramidase [Homo sapiens],3KEH_A Chain A, Glucocerebrosidase [Homo sapiens],3KEH_B Chain B, Glucocerebrosidase [Homo sapiens] |
| 2WKL_A | 5.08e-34 | 48 | 470 | 67 | 496 | Velaglucerasealfa [Homo sapiens],2WKL_B Velaglucerase alfa [Homo sapiens],5LVX_A Crystal structure of glucocerebrosidase with an inhibitory quinazoline modulator [Homo sapiens],5LVX_B Crystal structure of glucocerebrosidase with an inhibitory quinazoline modulator [Homo sapiens],5LVX_C Crystal structure of glucocerebrosidase with an inhibitory quinazoline modulator [Homo sapiens],5LVX_D Crystal structure of glucocerebrosidase with an inhibitory quinazoline modulator [Homo sapiens],6TJK_AAA Chain AAA, Lysosomal acid glucosylceramidase [Homo sapiens],6TJK_BBB Chain BBB, Lysosomal acid glucosylceramidase [Homo sapiens],6TJQ_BBB Chain BBB, Glucosylceramidase [Homo sapiens],6TN1_AAA Chain AAA, Lysosomal acid glucosylceramidase [Homo sapiens],6YTR_AAA Chain AAA, Lysosomal acid glucosylceramidase [Homo sapiens],6YTR_BBB Chain BBB, Lysosomal acid glucosylceramidase [Homo sapiens],6Z3I_BBB Chain BBB, Lysosomal acid glucosylceramidase [Homo sapiens],7NWV_AAA Chain AAA, Lysosomal acid glucosylceramidase [Homo sapiens],7NWV_BBB Chain BBB, Lysosomal acid glucosylceramidase [Homo sapiens] |
| 1OGS_A | 1.25e-33 | 48 | 468 | 67 | 494 | humanacid-beta-glucosidase [Homo sapiens],1OGS_B human acid-beta-glucosidase [Homo sapiens],1Y7V_A Chain A, Glucosylceramidase [Homo sapiens],1Y7V_B Chain B, Glucosylceramidase [Homo sapiens],2F61_A Crystal structure of partially deglycosylated acid beta-glucosidase [Homo sapiens],2F61_B Crystal structure of partially deglycosylated acid beta-glucosidase [Homo sapiens],2J25_A Partially deglycosylated glucoceramidase [Homo sapiens],2J25_B Partially deglycosylated glucoceramidase [Homo sapiens],2NSX_A Structure of acid-beta-glucosidase with pharmacological chaperone provides insight into Gaucher disease [Homo sapiens],2NSX_B Structure of acid-beta-glucosidase with pharmacological chaperone provides insight into Gaucher disease [Homo sapiens],2NSX_C Structure of acid-beta-glucosidase with pharmacological chaperone provides insight into Gaucher disease [Homo sapiens],2NSX_D Structure of acid-beta-glucosidase with pharmacological chaperone provides insight into Gaucher disease [Homo sapiens],2NT0_A Acid-beta-glucosidase low pH, glycerol bound [Homo sapiens],2NT0_B Acid-beta-glucosidase low pH, glycerol bound [Homo sapiens],2NT0_C Acid-beta-glucosidase low pH, glycerol bound [Homo sapiens],2NT0_D Acid-beta-glucosidase low pH, glycerol bound [Homo sapiens],2NT1_A Structure of acid-beta-glucosidase at neutral pH [Homo sapiens],2NT1_B Structure of acid-beta-glucosidase at neutral pH [Homo sapiens],2NT1_C Structure of acid-beta-glucosidase at neutral pH [Homo sapiens],2NT1_D Structure of acid-beta-glucosidase at neutral pH [Homo sapiens],3GXD_A Crystal structure of Apo acid-beta-glucosidase pH 4.5 [Homo sapiens],3GXD_B Crystal structure of Apo acid-beta-glucosidase pH 4.5 [Homo sapiens],3GXD_C Crystal structure of Apo acid-beta-glucosidase pH 4.5 [Homo sapiens],3GXD_D Crystal structure of Apo acid-beta-glucosidase pH 4.5 [Homo sapiens],3GXF_A Crystal structure of acid-beta-glucosidase with isofagomine at neutral pH [Homo sapiens],3GXF_B Crystal structure of acid-beta-glucosidase with isofagomine at neutral pH [Homo sapiens],3GXF_C Crystal structure of acid-beta-glucosidase with isofagomine at neutral pH [Homo sapiens],3GXF_D Crystal structure of acid-beta-glucosidase with isofagomine at neutral pH [Homo sapiens],3GXI_A Crystal structure of acid-beta-glucosidase at pH 5.5 [Homo sapiens],3GXI_B Crystal structure of acid-beta-glucosidase at pH 5.5 [Homo sapiens],3GXI_C Crystal structure of acid-beta-glucosidase at pH 5.5 [Homo sapiens],3GXI_D Crystal structure of acid-beta-glucosidase at pH 5.5 [Homo sapiens],3GXM_A Crystal structure of acid-beta-glucosidase at pH 4.5, phosphate crystallization condition [Homo sapiens],3GXM_B Crystal structure of acid-beta-glucosidase at pH 4.5, phosphate crystallization condition [Homo sapiens],3GXM_C Crystal structure of acid-beta-glucosidase at pH 4.5, phosphate crystallization condition [Homo sapiens],3GXM_D Crystal structure of acid-beta-glucosidase at pH 4.5, phosphate crystallization condition [Homo sapiens],3RIK_A The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIK_B The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIK_C The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIK_D The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIL_A The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIL_B The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIL_C The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],3RIL_D The acid beta-glucosidase active site exhibits plasticity in binding 3,4,5,6-tetrahydroxyazepane-based inhibitors: implications for pharmacological chaperone design for gaucher disease [Homo sapiens],6MOZ_A Structure of acid-beta-glucosidase in complex with an aromatic pyrrolidine iminosugar inhibitor [Homo sapiens],6MOZ_B Structure of acid-beta-glucosidase in complex with an aromatic pyrrolidine iminosugar inhibitor [Homo sapiens],6Q1N_A Glucocerebrosidase in complex with pharmacological chaperone IMX8 [Homo sapiens],6Q1N_B Glucocerebrosidase in complex with pharmacological chaperone IMX8 [Homo sapiens],6Q1P_A Glucocerebrosidase in complex with pharmacological chaperone norIMX8 [Homo sapiens],6Q1P_B Glucocerebrosidase in complex with pharmacological chaperone norIMX8 [Homo sapiens],6Q6K_A Crystal structure of recombinant human beta-glucocerebrosidase in complex with cyclophellitol activity based probe with Cy5 tag (ME569) [Homo sapiens],6Q6K_B Crystal structure of recombinant human beta-glucocerebrosidase in complex with cyclophellitol activity based probe with Cy5 tag (ME569) [Homo sapiens],6Q6L_A Crystal structure of recombinant human beta-glucocerebrosidase in complex with adamantyl-cyclophellitol inhibitor (ME656) [Homo sapiens],6Q6L_B Crystal structure of recombinant human beta-glucocerebrosidase in complex with adamantyl-cyclophellitol inhibitor (ME656) [Homo sapiens],6Q6N_A Crystal structure of recombinant human beta-glucocerebrosidase in complex with biphenyl-cyclophellitol inhibitor (ME655) [Homo sapiens],6Q6N_B Crystal structure of recombinant human beta-glucocerebrosidase in complex with biphenyl-cyclophellitol inhibitor (ME655) [Homo sapiens],6TJJ_AAA Chain AAA, Glucosylceramidase [Homo sapiens],6TJJ_BBB Chain BBB, Glucosylceramidase [Homo sapiens],6YTP_AAA Chain AAA, Glucosylceramidase [Homo sapiens],6YTP_BBB Chain BBB, Glucosylceramidase [Homo sapiens],6YUT_AAA Chain AAA, Glucosylceramidase [Homo sapiens],6YUT_BBB Chain BBB, Glucosylceramidase [Homo sapiens],6YV3_AAA Chain AAA, Glucosylceramidase [Homo sapiens],6YV3_BBB Chain BBB, Glucosylceramidase [Homo sapiens],6Z39_AAA Chain AAA, Glucosylceramidase [Homo sapiens],6Z39_BBB Chain BBB, Glucosylceramidase [Homo sapiens] |
Swiss-Prot Hits download full data without filtering help
| Hit ID | E-Value | Query Start | Query End | Hit Start | Hit End | Description |
|---|---|---|---|---|---|---|
| O16580 | 2.59e-37 | 48 | 437 | 87 | 481 | Putative glucosylceramidase 1 OS=Caenorhabditis elegans OX=6239 GN=gba-1 PE=1 SV=2 |
| Q9UB00 | 4.50e-37 | 14 | 469 | 52 | 519 | Putative glucosylceramidase 4 OS=Caenorhabditis elegans OX=6239 GN=gba-4 PE=3 SV=2 |
| G5ECR8 | 9.57e-36 | 33 | 470 | 77 | 520 | Putative glucosylceramidase 3 OS=Caenorhabditis elegans OX=6239 GN=gba-3 PE=3 SV=1 |
| Q2KHZ8 | 2.46e-33 | 48 | 470 | 106 | 535 | Lysosomal acid glucosylceramidase OS=Bos taurus OX=9913 GN=GBA PE=2 SV=1 |
| Q9BDT0 | 4.45e-33 | 48 | 470 | 106 | 535 | Lysosomal acid glucosylceramidase OS=Pan troglodytes OX=9598 GN=GBA PE=3 SV=1 |
SignalP and Lipop Annotations help
This protein is predicted as SP
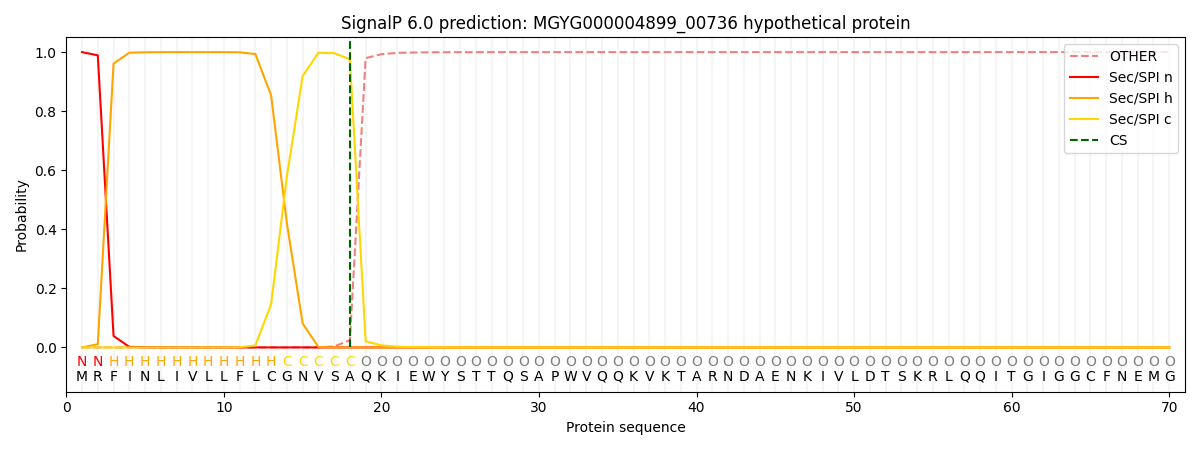
| Other | SP_Sec_SPI | LIPO_Sec_SPII | TAT_Tat_SPI | TATLIP_Sec_SPII | PILIN_Sec_SPIII |
|---|---|---|---|---|---|
| 0.000276 | 0.999091 | 0.000202 | 0.000140 | 0.000129 | 0.000129 |
